Protein Visualization in VSCode
VSCode Python add-in now supports ipywidgets in Jupyter Notebooks so you can view a protein or molecule visualization directly within the VSCode environment.
First, we need to load the nglview library into the Jupyter Notebook environment. To do this, I am going to use anaconda.
Quick note, the first time I tried to install the packages, I got several errors; however, after updating to the latest stable versions of anaconda and node, everything ran a lot smoother.
I am using the following versions of anaconda and node:
anaconda 4.8.3
node v12.16.2
Open the Anaconda command line and type the following command to install the nglview packages:
conda install nglviewOnce the nglview packages have downloaded and installed, install the mdanalysis stable release using the following commands:
conda config --add channels conda-forge
conda install mdanalysisI also installed the mdanalysis tests using the following command:
conda install MDAnalysisTestsFinally, install the ipywidgets using the following command:
conda install ipywidgetsOnce the ipywidgets have installed, we start the Jupyter Notebook environment, open an existing or new notebook, and run the following command to visualize the protein.
import MDAnalysis as mda
import nglview as nv
from nglview.datafiles import PDB, XTC
u = mda.Universe(PDB, XTC)
protein = u.select_atoms('protein')
w = nv.show_mdanalysis(protein)
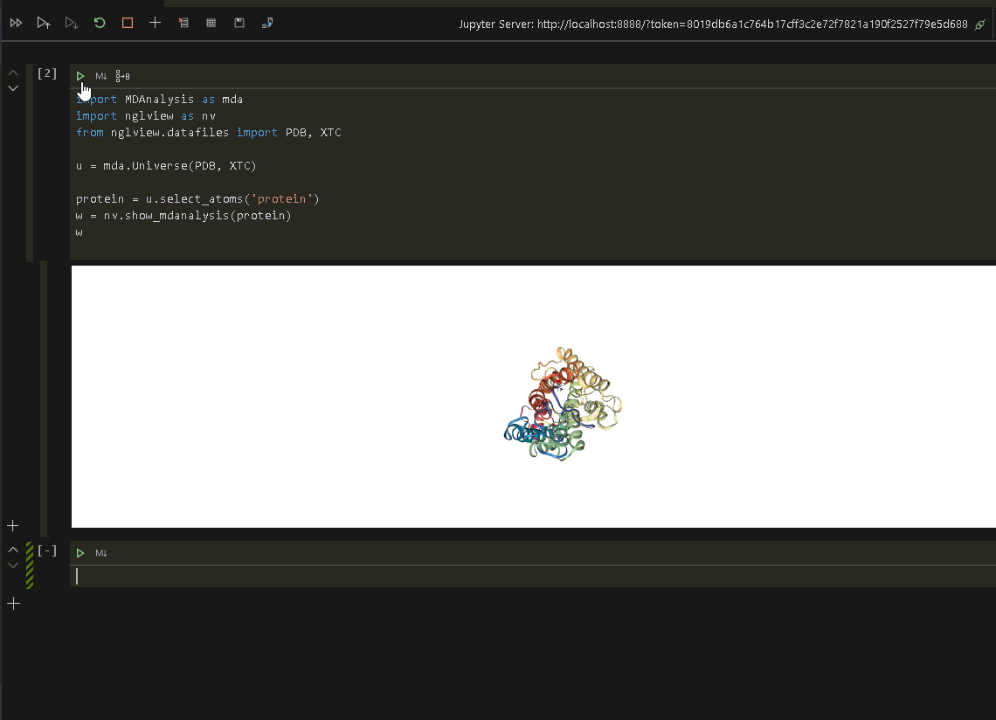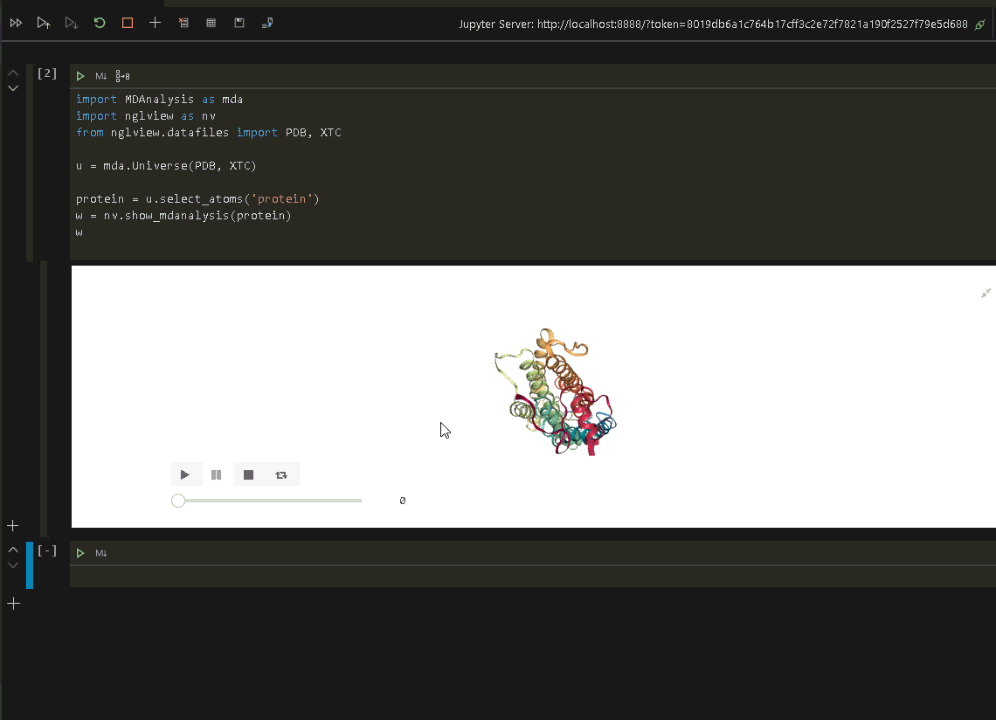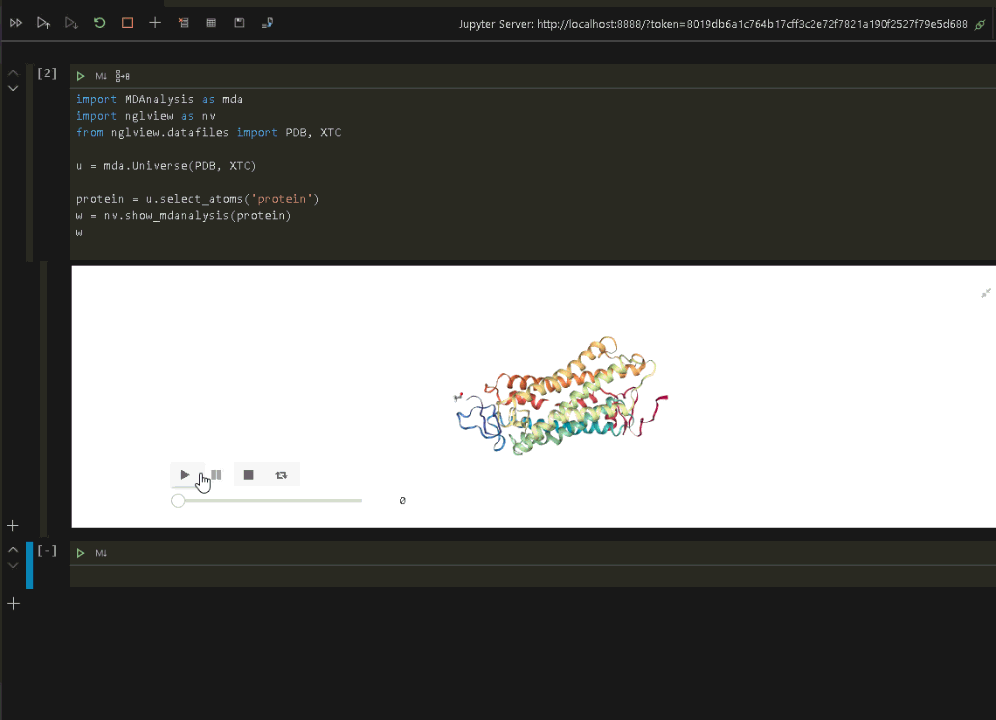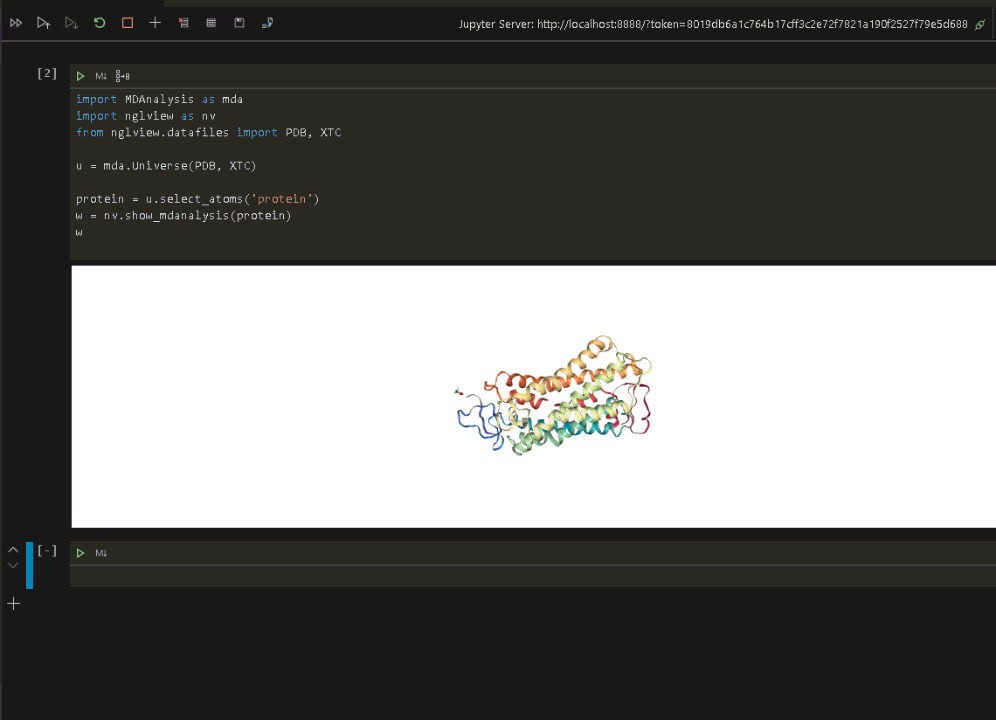
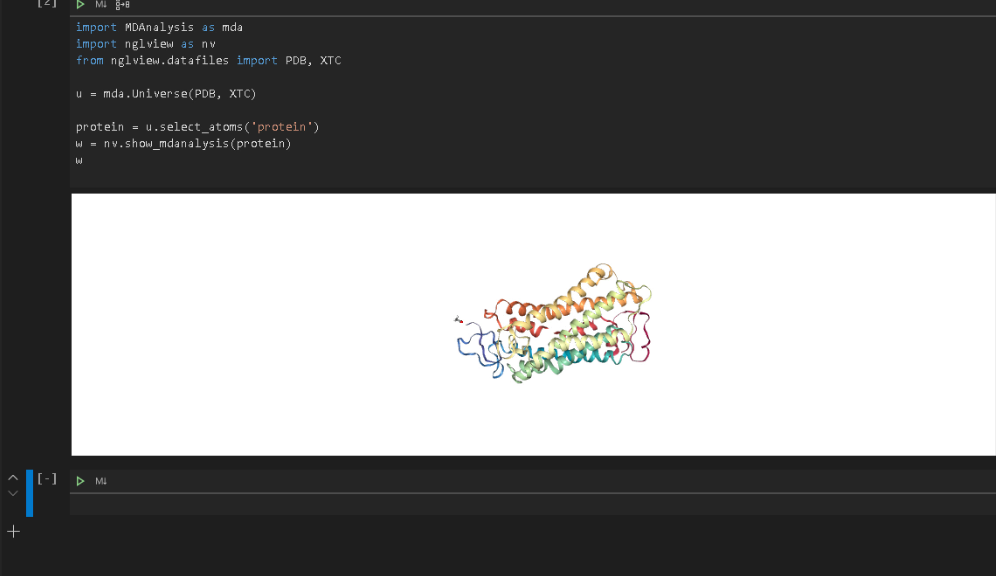
wRun the code to display the protein in VSCode.
Now that we have verified that everything is up and running in the Jupyter environment, we can connect VSCode to the same Jupyter environment and run the visualization (see the previous post on connecting VSCode to a Jupyter Notebook).
Once the Jupyter connection is made, and a notebook opened, we can rerun the protein visualization from within VSCode.